PP-D100T Pichia Pastoris Host Cell DNA Residue Detection kit
The Pichia Pastoris Host Cell DNA Residue Detection Kit can be used for Quantitative analysis of DNA residue in recombinant protein expressed products, purified intermediate and finished products from host cell..

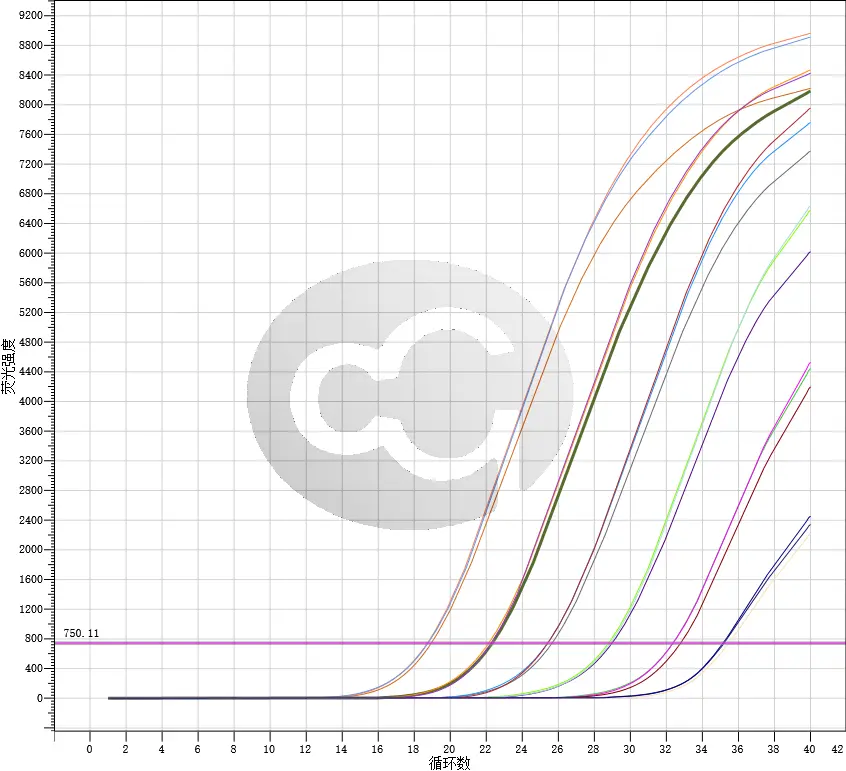
.webp)